Discover indicators
Let’s set our health atlas. For this example we will use the Chicago
Health Atlas. We can do so by calling ha_set() with the
Chicago Health Atlas URL.
ha_set("chicagohealthatlas.org")If we need to check which health atlas we are using, we can use
ha_get().
ha_get()
#> [1] "https://chicagohealthatlas.org/api/v1/"We can list all the topics (aka indicators) present within Chicago
Health Atlas by using ha_topics(). The most important
column here is the topic_key. An individual
topic_key can be used to identify a topic within subsequent
functions.
topics <- ha_topics(progress = FALSE)
#> Error in get(paste0(generic, ".", class), envir = get_method_env()) :
#> object 'type_sum.accel' not found
topics
#> # A tibble: 409 × 7
#> topic_name topic_key topic_description topic_units subcategory_name
#> <chr> <chr> <chr> <chr> <chr>
#> 1 9th grade education… EDA Residents 25 or … % of resid… Education
#> 2 ACA marketplace enr… ENR Number of plan s… plan selec… Access to Care
#> 3 Accidents mortality VRAC Number of people… count of d… Injury & Violen…
#> 4 Accidents mortality… VRACR Age-adjusted rat… per 100,00… Injury & Violen…
#> 5 Active business lic… CHANVYI Count of active … licenses p… Income
#> 6 Adult asthma HCSATH Number of adults… count of a… Chronic Disease
#> 7 Adult asthma rate HCSATHP Percent of adult… % of adults Chronic Disease
#> 8 Adult binge drinking HCSBD Number of adults… count of a… Alcohol & Drug …
#> 9 Adult binge drinkin… HCSBDP Percent of adult… % of adults Alcohol & Drug …
#> 10 Adult diabetes HCSDIA Number of adults… count of a… Chronic Disease
#> # ℹ 399 more rows
#> # ℹ 2 more variables: subcategory_key <chr>, category_name <chr>There may be a specific topic area you are interested in exploring.
You can explore these topic areas using
ha_subcategories().
subcategories <- ha_subcategories()
subcategories
#> # A tibble: 30 × 3
#> subcategory_name subcategory_key category_name
#> <chr> <chr> <chr>
#> 1 Access to Care access-to-care Clinical Care
#> 2 Quality of Care quality-of-care Clinical Care
#> 3 Community Safety community-safety-1 Physical Environment
#> 4 Housing & Transit housing-transit Physical Environment
#> 5 Pollution pollution Physical Environment
#> 6 Resource Availability resource-availability Physical Environment
#> 7 Behavioral Health behavioral-health Morbidity
#> 8 Chronic Disease chronic-disease-1 Morbidity
#> 9 Infectious Disease infectious-disease-1 Morbidity
#> 10 Injury & Violence injury-violence-1 Morbidity
#> # ℹ 20 more rowsYou can use a subcategory_key to subset the list of
topics.
ha_topics("diet-exercise")
#> # A tibble: 20 × 7
#> topic_name topic_key topic_description topic_units subcategory_name
#> <chr> <chr> <chr> <chr> <chr>
#> 1 Adult fruit and veg… HCSFV "Number of adult… count of a… Diet & Exercise
#> 2 Adult fruit and veg… HCSFVP "Percent of adul… % of adults Diet & Exercise
#> 3 Adult physical inac… HCSPA "Number of adult… count of a… Diet & Exercise
#> 4 Adult physical inac… HCSPAP "Percent of adul… % of adults Diet & Exercise
#> 5 Adult soda consumpt… HCSS "Number of adult… count of a… Diet & Exercise
#> 6 Adult soda consumpt… HCSSP "Percent of adul… % of adults Diet & Exercise
#> 7 Easy access to frui… HCSFVA "Number of adult… count of a… Diet & Exercise
#> 8 Easy access to frui… HCSFVAP "Percent of adul… % of adults Diet & Exercise
#> 9 High School fruit a… YRFV "Number of Chica… count of s… Diet & Exercise
#> 10 High School fruit a… YRFVP "Percent of Chic… % of stude… Diet & Exercise
#> 11 High School physica… YRPA "Number of Chica… count of s… Diet & Exercise
#> 12 High School physica… YRPAP "Percent of Chic… % of stude… Diet & Exercise
#> 13 High School physica… YRPI "Number of Chica… count of s… Diet & Exercise
#> 14 High School physica… YRPIP "Percent of Chic… % of stude… Diet & Exercise
#> 15 High School soda co… YRSO "Number of Chica… count of s… Diet & Exercise
#> 16 High School soda co… YRSOP "Percent of Chic… % of stude… Diet & Exercise
#> 17 Middle School physi… YRMPA "Number of Chica… count of s… Diet & Exercise
#> 18 Middle School physi… YRMPAP "Percent of Chic… % of stude… Diet & Exercise
#> 19 Middle School physi… YRMPI "Number of Chica… count of s… Diet & Exercise
#> 20 Middle School physi… YRMPIP "Percent of Chic… % of stude… Diet & Exercise
#> # ℹ 2 more variables: subcategory_key <chr>, category_name <chr>Once we have a topic or topics in mind, we can explore what
populations, time periods, and geographic scales that data is available
at by using ha_coverage(). Again, the most important
columns here are the key columns which can be used to specify the data
desired.
coverage <- ha_coverage("HCSFVAP", progress = FALSE)
coverage
#> # A tibble: 166 × 7
#> topic_key population_key population_name population_grouping period_key
#> <chr> <chr> <chr> <chr> <chr>
#> 1 HCSFVAP "" Full population "" 2020-2021
#> 2 HCSFVAP "" Full population "" 2016-2018
#> 3 HCSFVAP "" Full population "" 2015-2017
#> 4 HCSFVAP "" Full population "" 2014-2016
#> 5 HCSFVAP "" Full population "" 2022-2023
#> 6 HCSFVAP "" Full population "" 2021-2022
#> 7 HCSFVAP "" Full population "" 2022-2023
#> 8 HCSFVAP "" Full population "" 2023
#> 9 HCSFVAP "" Full population "" 2021-2022
#> 10 HCSFVAP "" Full population "" 2022
#> # ℹ 156 more rows
#> # ℹ 2 more variables: layer_key <chr>, layer_name <chr>Import tabular data
Now, we can import our data using ha_data() and
specifying the keys we identified above.
ease_of_access <- ha_data(
topic_key = "HCSFVAP",
population_key = "",
period_key = "2022-2023",
layer_key = "neighborhood"
)
ease_of_access
#> # A tibble: 77 × 7
#> geoid topic_key population_key period_key layer_key value standardError
#> <chr> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1714000-35 HCSFVAP "" 2022-2023 neighborh… 57.9 6.86
#> 2 1714000-36 HCSFVAP "" 2022-2023 neighborh… 54.7 6.25
#> 3 1714000-37 HCSFVAP "" 2022-2023 neighborh… 45.5 7.45
#> 4 1714000-38 HCSFVAP "" 2022-2023 neighborh… 56.9 5.86
#> 5 1714000-39 HCSFVAP "" 2022-2023 neighborh… 52.4 10.5
#> 6 1714000-4 HCSFVAP "" 2022-2023 neighborh… 71.7 5.49
#> 7 1714000-40 HCSFVAP "" 2022-2023 neighborh… 36.8 6.73
#> 8 1714000-41 HCSFVAP "" 2022-2023 neighborh… 65.9 7.52
#> 9 1714000-42 HCSFVAP "" 2022-2023 neighborh… 48.4 8.27
#> 10 1714000-1 HCSFVAP "" 2022-2023 neighborh… 56.7 4.96
#> # ℹ 67 more rowsWe can even specify multiple topics, populations, and periods to get
data for. ha_data() will return a combined table with data
for every combination of topic, population, and period requested. A
warning will be given for every invalid combindation of topic,
population, and period requested.
combinations_of_data <- ha_data(
topic_key = c("POP", "UMP"),
population_key = c("", "H"),
period_key = c("2017-2021", "2018-2022", "invalid"),
layer_key = "neighborhood"
)
#> Warning: Your API call has errors. No results for topic_key = "POP"
#> population_key = "" period_key = "invalid" layer_key = "neighborhood".
#> Warning: Your API call has errors. No results for topic_key = "UMP"
#> population_key = "" period_key = "invalid" layer_key = "neighborhood".
#> Warning: Your API call has errors. No results for topic_key = "POP"
#> population_key = "H" period_key = "invalid" layer_key = "neighborhood".
#> Warning: Your API call has errors. No results for topic_key = "UMP"
#> population_key = "H" period_key = "invalid" layer_key = "neighborhood".
combinations_of_data
#> # A tibble: 616 × 7
#> geoid topic_key population_key period_key layer_key value standardError
#> <chr> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1714000-35 POP "" 2017-2021 neighbor… 21276. NA
#> 2 1714000-36 POP "" 2017-2021 neighbor… 7417. NA
#> 3 1714000-37 POP "" 2017-2021 neighbor… 2280. NA
#> 4 1714000-38 POP "" 2017-2021 neighbor… 24397. NA
#> 5 1714000-39 POP "" 2017-2021 neighbor… 18503. NA
#> 6 1714000-4 POP "" 2017-2021 neighbor… 42252. NA
#> 7 1714000-40 POP "" 2017-2021 neighbor… 11608. NA
#> 8 1714000-41 POP "" 2017-2021 neighbor… 28802. NA
#> 9 1714000-42 POP "" 2017-2021 neighbor… 24362. NA
#> 10 1714000-1 POP "" 2017-2021 neighbor… 55627. NA
#> # ℹ 606 more rowsIf you want to mix and match topics, populations, years, or layers of
data, I recommend creating a table of all the datasets you want, and
purrr::pmap()-ing over the table.
library(tibble)
library(purrr)
# creating a table of data I want
metadata <- tribble(
~ topic_key, ~ population_key, ~ period_key, ~ layer_key,
"POP", "", "2017-2021", "neighborhood",
"HCSFVAP", "", "2020-2021", "neighborhood",
"UMP", "H", "2017-2021", "neighborhood",
)
metadata %>%
pmap(ha_data)
#> [[1]]
#> # A tibble: 77 × 7
#> geoid topic_key population_key period_key layer_key value standardError
#> <chr> <chr> <chr> <chr> <chr> <dbl> <lgl>
#> 1 1714000-35 POP "" 2017-2021 neighbor… 21276. NA
#> 2 1714000-36 POP "" 2017-2021 neighbor… 7417. NA
#> 3 1714000-37 POP "" 2017-2021 neighbor… 2280. NA
#> 4 1714000-38 POP "" 2017-2021 neighbor… 24397. NA
#> 5 1714000-39 POP "" 2017-2021 neighbor… 18503. NA
#> 6 1714000-4 POP "" 2017-2021 neighbor… 42252. NA
#> 7 1714000-40 POP "" 2017-2021 neighbor… 11608. NA
#> 8 1714000-41 POP "" 2017-2021 neighbor… 28802. NA
#> 9 1714000-42 POP "" 2017-2021 neighbor… 24362. NA
#> 10 1714000-1 POP "" 2017-2021 neighbor… 55627. NA
#> # ℹ 67 more rows
#>
#> [[2]]
#> # A tibble: 77 × 7
#> geoid topic_key population_key period_key layer_key value standardError
#> <chr> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1714000-35 HCSFVAP "" 2020-2021 neighborh… 52.0 9.18
#> 2 1714000-36 HCSFVAP "" 2020-2021 neighborh… 63.8 9.34
#> 3 1714000-37 HCSFVAP "" 2020-2021 neighborh… 33.2 9.78
#> 4 1714000-38 HCSFVAP "" 2020-2021 neighborh… 47.4 7.36
#> 5 1714000-39 HCSFVAP "" 2020-2021 neighborh… 61.3 7.22
#> 6 1714000-4 HCSFVAP "" 2020-2021 neighborh… 77.4 5.22
#> 7 1714000-40 HCSFVAP "" 2020-2021 neighborh… 46.2 8.67
#> 8 1714000-41 HCSFVAP "" 2020-2021 neighborh… 76.3 5.30
#> 9 1714000-42 HCSFVAP "" 2020-2021 neighborh… 56.4 7.77
#> 10 1714000-1 HCSFVAP "" 2020-2021 neighborh… 58.0 4.98
#> # ℹ 67 more rows
#>
#> [[3]]
#> # A tibble: 77 × 7
#> geoid topic_key population_key period_key layer_key value standardError
#> <chr> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 1714000-… UMP H 2017-2021 neighbor… 20.2 22.8
#> 2 1714000-… UMP H 2017-2021 neighbor… 0.0239 48.0
#> 3 1714000-… UMP H 2017-2021 neighbor… 6.60 21.8
#> 4 1714000-… UMP H 2017-2021 neighbor… 9.47 30.5
#> 5 1714000-… UMP H 2017-2021 neighbor… 12.2 28.4
#> 6 1714000-4 UMP H 2017-2021 neighbor… 8.13 3.35
#> 7 1714000-… UMP H 2017-2021 neighbor… 29.8 52.9
#> 8 1714000-… UMP H 2017-2021 neighbor… 6.42 10.5
#> 9 1714000-… UMP H 2017-2021 neighbor… 3.37 37.6
#> 10 1714000-1 UMP H 2017-2021 neighbor… 5.19 4.38
#> # ℹ 67 more rowsImport spatial data
We can see all the geographic layers available by using
ha_layers().
layers <- ha_layers()
layers
#> # A tibble: 4 × 4
#> layer_name layer_key layer_description layer_url
#> <chr> <chr> <chr> <chr>
#> 1 Community areas neighborhood The city of Chicago is divided into 77… https://…
#> 2 ZIP Codes zip The ZIP code is a basic unit of geogra… https://…
#> 3 Census Tracts tract-2020 Census tracts are small geographies de… https://…
#> 4 Chicago place Cities, towns, villages, and boroughs,… https://…Since we just downloaded our data at the Community Area level, let’s
import the Community Area geographic layer with
ha_layer().
community_areas <- ha_layer("neighborhood")
community_areas
#> Simple feature collection with 77 features and 6 fields
#> Geometry type: MULTIPOLYGON
#> Dimension: XY
#> Bounding box: xmin: -87.94011 ymin: 41.64454 xmax: -87.52419 ymax: 42.02305
#> Geodetic CRS: WGS 84
#> First 10 features:
#> geoid layer_key name population state
#> 1 1714000-1 neighborhood Rogers Park (Chicago, IL) 55454 IL
#> 2 1714000-10 neighborhood Norwood Park (Chicago, IL) 41069 IL
#> 3 1714000-11 neighborhood Jefferson Park (Chicago, IL) 26201 IL
#> 4 1714000-12 neighborhood Forest Glen (Chicago, IL) 19579 IL
#> 5 1714000-13 neighborhood North Park (Chicago, IL) 17522 IL
#> 6 1714000-14 neighborhood Albany Park (Chicago, IL) 48549 IL
#> 7 1714000-15 neighborhood Portage Park (Chicago, IL) 63038 IL
#> 8 1714000-16 neighborhood Irving Park (Chicago, IL) 51911 IL
#> 9 1714000-17 neighborhood Dunning (Chicago, IL) 43120 IL
#> 10 1714000-18 neighborhood Montclare (Chicago, IL) 14412 IL
#> notes geometry
#> 1 Far North Side MULTIPOLYGON (((-87.65456 4...
#> 2 Far North Side MULTIPOLYGON (((-87.78002 4...
#> 3 Far North Side MULTIPOLYGON (((-87.75264 4...
#> 4 Far North Side MULTIPOLYGON (((-87.72642 4...
#> 5 Far North Side MULTIPOLYGON (((-87.7069 41...
#> 6 Far North Side MULTIPOLYGON (((-87.70404 4...
#> 7 Northwest Side MULTIPOLYGON (((-87.75264 4...
#> 8 Northwest Side MULTIPOLYGON (((-87.69475 4...
#> 9 Northwest Side MULTIPOLYGON (((-87.77621 4...
#> 10 Northwest Side MULTIPOLYGON (((-87.78942 4...You can also set geometry = TRUE within your data call
to get the geographic layer’s geometry along with your data.
ease_of_access <- ha_data(
topic_key = "HCSFVAP",
population_key = "",
period_key = "2022-2023",
layer_key = "neighborhood",
geometry = TRUE
)
ease_of_access
#> Simple feature collection with 77 features and 7 fields
#> Geometry type: MULTIPOLYGON
#> Dimension: XY
#> Bounding box: xmin: -87.94011 ymin: 41.64454 xmax: -87.52419 ymax: 42.02305
#> Geodetic CRS: WGS 84
#> First 10 features:
#> geoid topic_key population_key period_key layer_key value
#> 1 1714000-1 HCSFVAP 2022-2023 neighborhood 56.70447
#> 2 1714000-10 HCSFVAP 2022-2023 neighborhood 61.06724
#> 3 1714000-11 HCSFVAP 2022-2023 neighborhood 61.46267
#> 4 1714000-12 HCSFVAP 2022-2023 neighborhood 81.03884
#> 5 1714000-13 HCSFVAP 2022-2023 neighborhood 54.84689
#> 6 1714000-14 HCSFVAP 2022-2023 neighborhood 52.98553
#> 7 1714000-15 HCSFVAP 2022-2023 neighborhood 61.05424
#> 8 1714000-16 HCSFVAP 2022-2023 neighborhood 61.62744
#> 9 1714000-17 HCSFVAP 2022-2023 neighborhood 72.73395
#> 10 1714000-18 HCSFVAP 2022-2023 neighborhood 51.01435
#> standardError geometry
#> 1 4.958576 MULTIPOLYGON (((-87.65456 4...
#> 2 5.929492 MULTIPOLYGON (((-87.78002 4...
#> 3 5.845823 MULTIPOLYGON (((-87.75264 4...
#> 4 4.560229 MULTIPOLYGON (((-87.72642 4...
#> 5 10.003305 MULTIPOLYGON (((-87.7069 41...
#> 6 6.182114 MULTIPOLYGON (((-87.70404 4...
#> 7 5.687155 MULTIPOLYGON (((-87.75264 4...
#> 8 6.953888 MULTIPOLYGON (((-87.69475 4...
#> 9 5.353022 MULTIPOLYGON (((-87.77621 4...
#> 10 9.557330 MULTIPOLYGON (((-87.78942 4...Let’s map our data!
library(ggplot2)
plot <- ggplot(ease_of_access) +
geom_sf(aes(fill = value), alpha = 0.7) +
scale_fill_distiller(palette = "GnBu", direction = 1) +
labs(
title = "Easy Access to Fruits and Vegetables within Chicago",
fill = "Percent of adults who reported\nthat it is very easy for them to\nget fresh fruits and vegetables."
) +
theme_minimal()
plot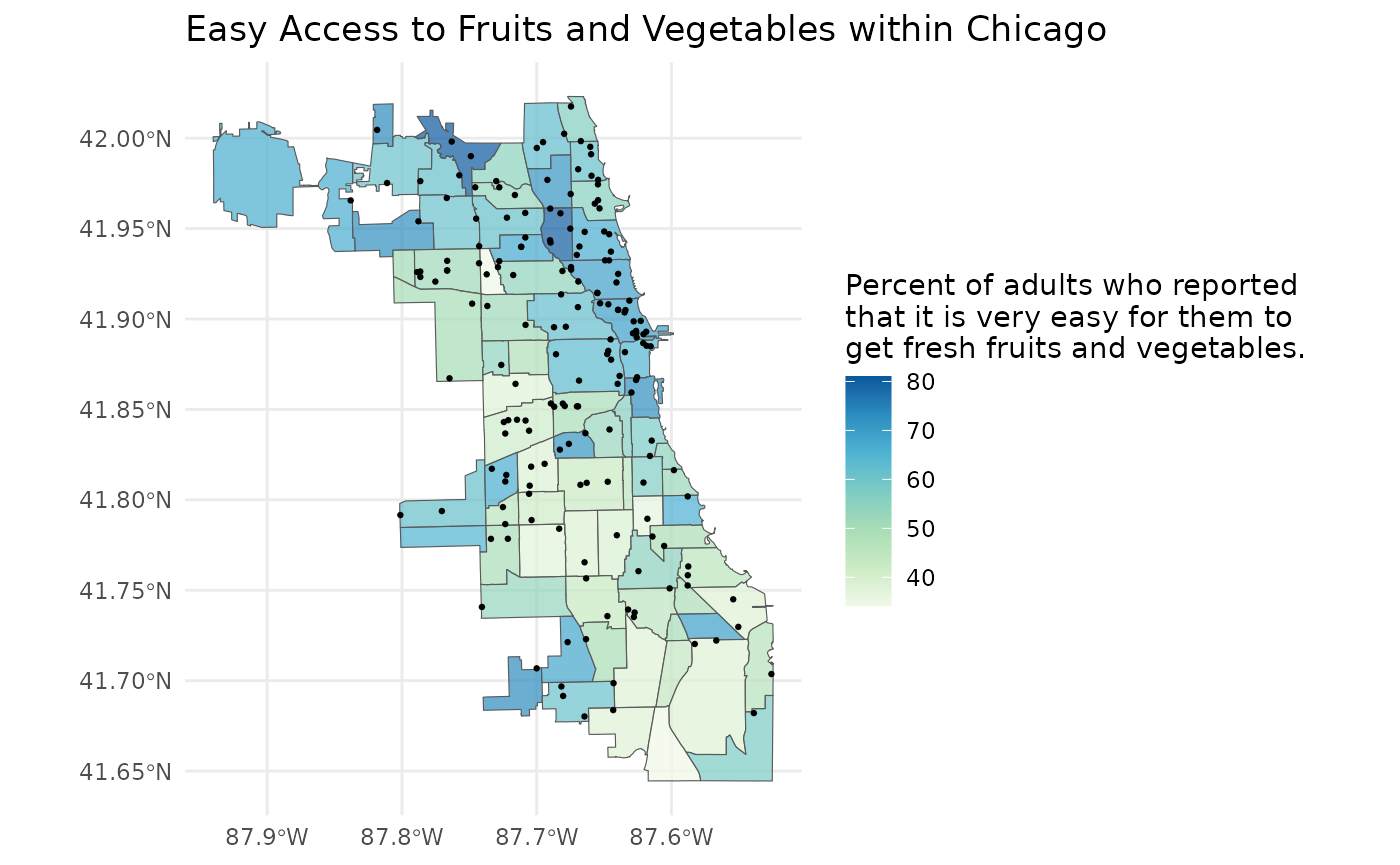
Our map looks pretty good, but perhaps there is a point layer that
may provide more insight into the spatial variation of the ease of
access to fruits and vegetables. We can use
ha_point_layers() to list all the point layers available in
the Chicago Health Atlas.
point_layers <- ha_point_layers()
point_layers
#> # A tibble: 10 × 3
#> point_layer_name point_layer_uuid point_layer_descript…¹
#> <chr> <chr> <chr>
#> 1 Acute Care Hospitals - 2023 67f58fa0-0dfa-4… ""
#> 2 Chicago Public Schools - 2023 5a449804-a2cc-4… ""
#> 3 Federally Qualified Health Centers -… 22f48fd6-ee98-4… ""
#> 4 Federally Qualified Health Centers (… f224b3ce-6d83-4… ""
#> 5 Grocery Stores 7d9caf3c-75e6-4… "All chain grocery st…
#> 6 Hospitals 8768fad7-65a2-4… "https://hifld-geopla…
#> 7 Nursing Homes 379a55c7-e569-4… "https://hifld-geopla…
#> 8 Pharmacies and Drug Stores 93ace519-6ba2-4… "All chain pharmacies…
#> 9 Skilled Nursing Facilities - 2023 93bc497d-3881-4… ""
#> 10 WIC Offices - 2023 7c8e9992-4e25-4… ""
#> # ℹ abbreviated name: ¹point_layer_descriptionGrocery store locations may be an important aspect of the ease of
access to fruits and vegetables. We can import this layer by providing
the point_layer_uuid to ha_point_layer().
grocery_stores <- ha_point_layer("7d9caf3c-75e6-4382-8c97-069696a3efbf")Now that we have imported our grocery stores, let’s layer them on top of our map.
plot +
geom_sf(data = grocery_stores, size = 0.5)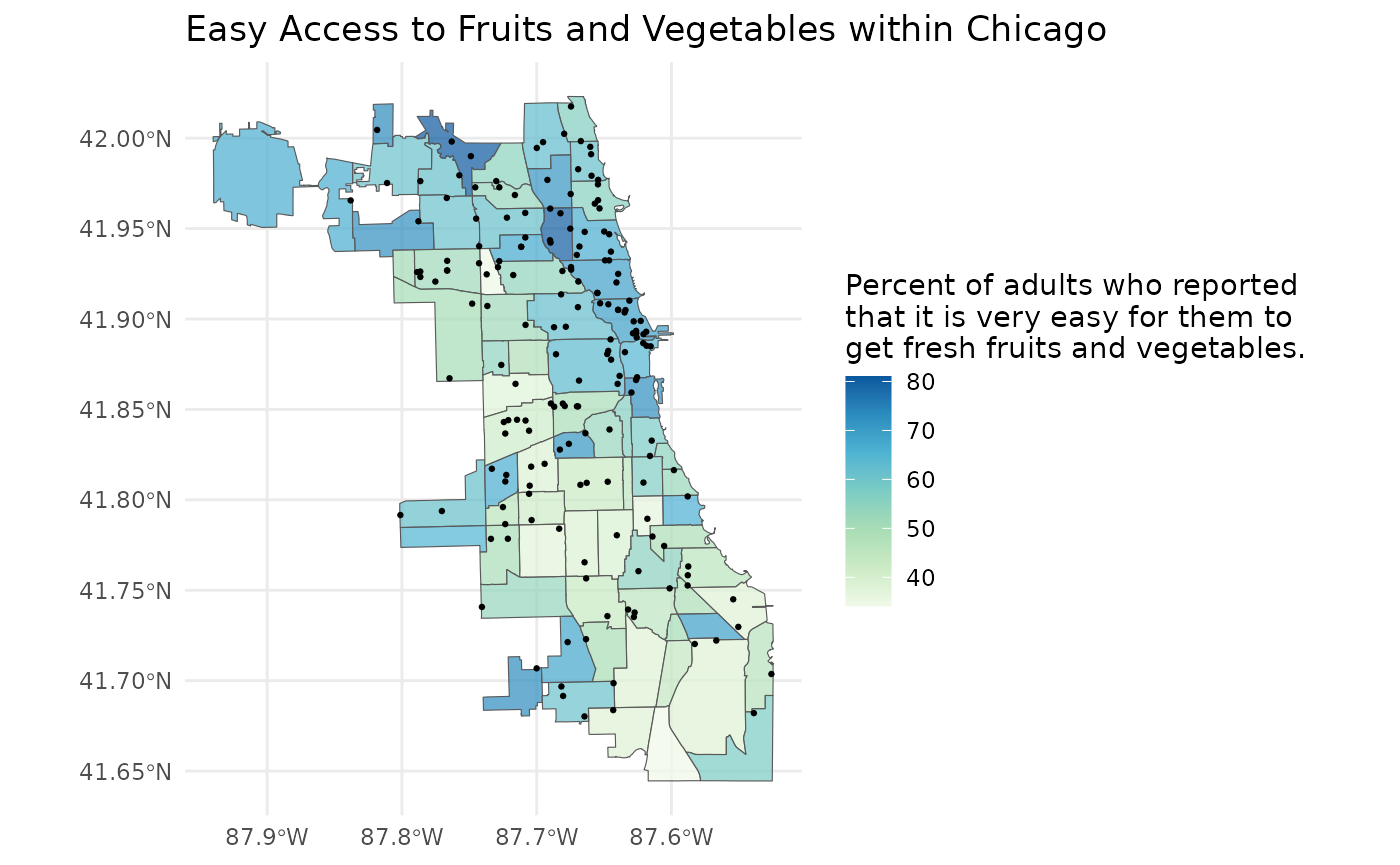
As expected, it seems that the areas with more grocery stores tend to have a higher percent of adults who report that it is very easy to get fresh fruits and vegetables.
This is a typical use case for the healthatlas in which
we explored every function that healthatlas has to offer.
Now it’s time for you to explore!
